Amey PasarkarI am a PhD student at Princeton University, working in Vertaix on efficient data curation and training algorithms for AI models. I am fortunate to be advised by Professor Adji Bousso Dieng and am generously supported by a NSF Graduate Research Fellowship. Prior to joining Princeton, I graduated from Columbia University with a degree in Operations Research, where I had the privilege of working under Professor Itsik Pe'er on designing probabalistic graphical models for microbial dynamics. I have also recently interned at Amazon Science on the Fire TV team. You can contact me at amey [dot] pasarkar [at] princeton [dot] edu. CV / Google Scholar / GitHub / LinkedIn |

|
ResearchI have a particular interest in generative modelling and using ML to enable large-scale dataset curation. I like thinking about these problems in the context of efficient training on image and text corpuses, as well as accelerating material discovery. |
|
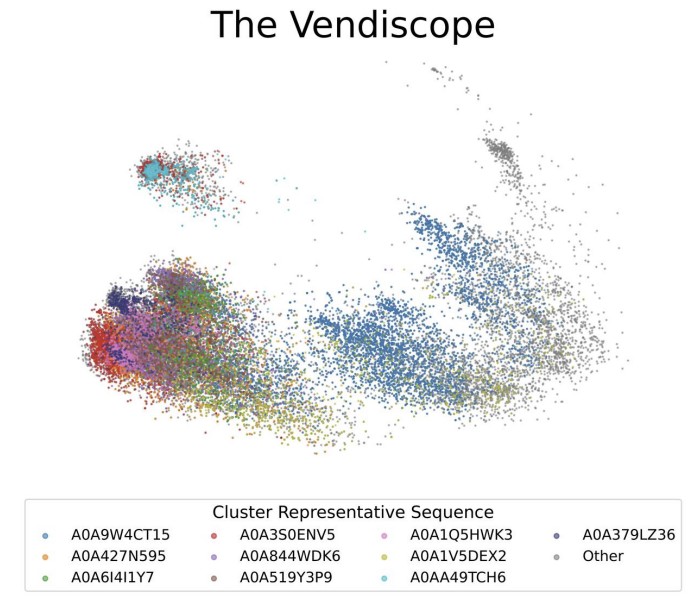
The Vendiscope: An Algorithmic Microscope For Data Collections
Amey P. Pasarkar, Adji Bousso Dieng arXiv pre-print, 2025 We introduce the Vendiscope, a scalable algorithmic microscope for analyzing the contents of datasets and behaviors of models in any domain where similarity can be defined. The Vendiscope can detect outliers and duplicates, diagnose model failure modes, and reveal memorization patterns across modalities, all at scale. |
|
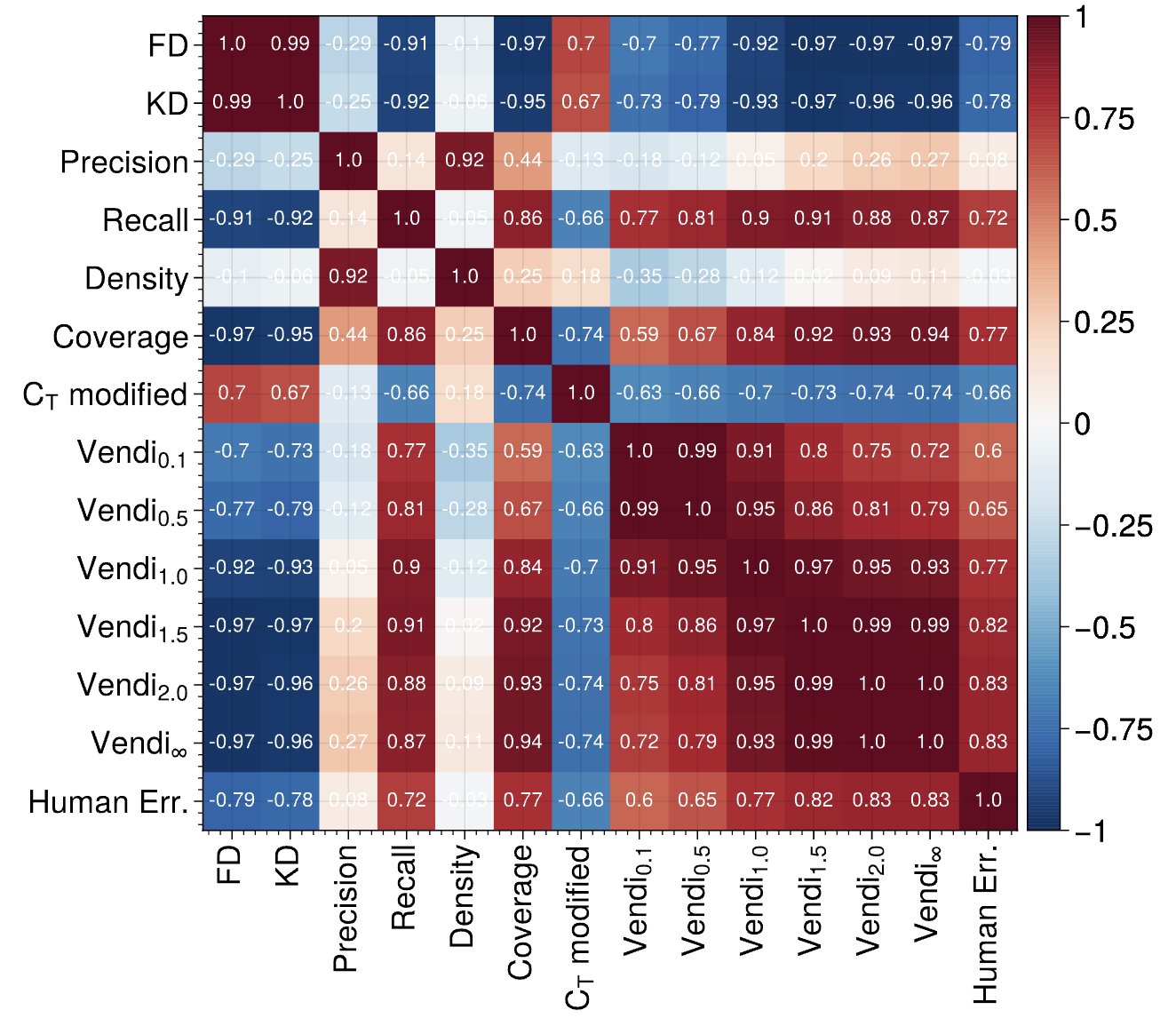
Cousins Of The Vendi Score: A Family Of Similarity-Based Diversity Metrics For Science And Machine Learning
Amey P. Pasarkar, Adji Bousso Dieng AISTATS, 2024 We generalize the Vendi Score to an entire family of metrics called the "Vendi scores". We demonstrate the utility of these metrics in evaluating image generative models, uncovering patterns of model memorization. |
|
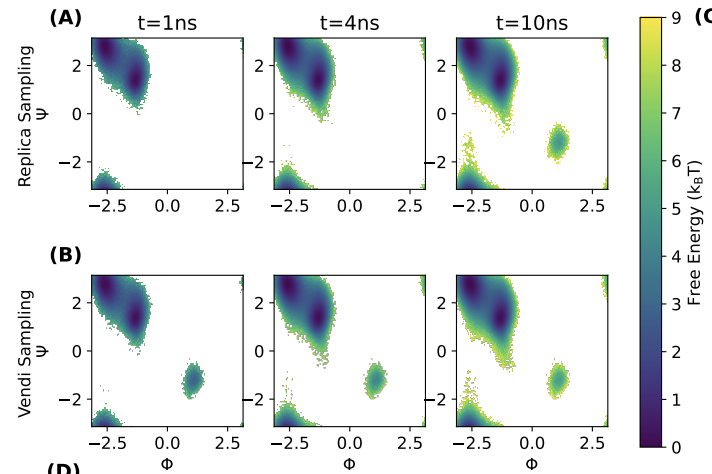
Vendi Sampling For Molecular Simulations: Diversity As A Force For Faster Convergence And Better Exploration
Amey P. Pasarkar, Gianluca M. Bencomo, Simon Olsson, Adji Bousso Dieng Journal of Chemical Physics, 2023 We propose Vendi Sampling, a method for fast sampling from complex Boltzmann distributions that encourages diversity throughout the sampling process. We demonstrate on various molecular systems that Vendi Sampling outperforms standard Langevin Dynamics. |
|
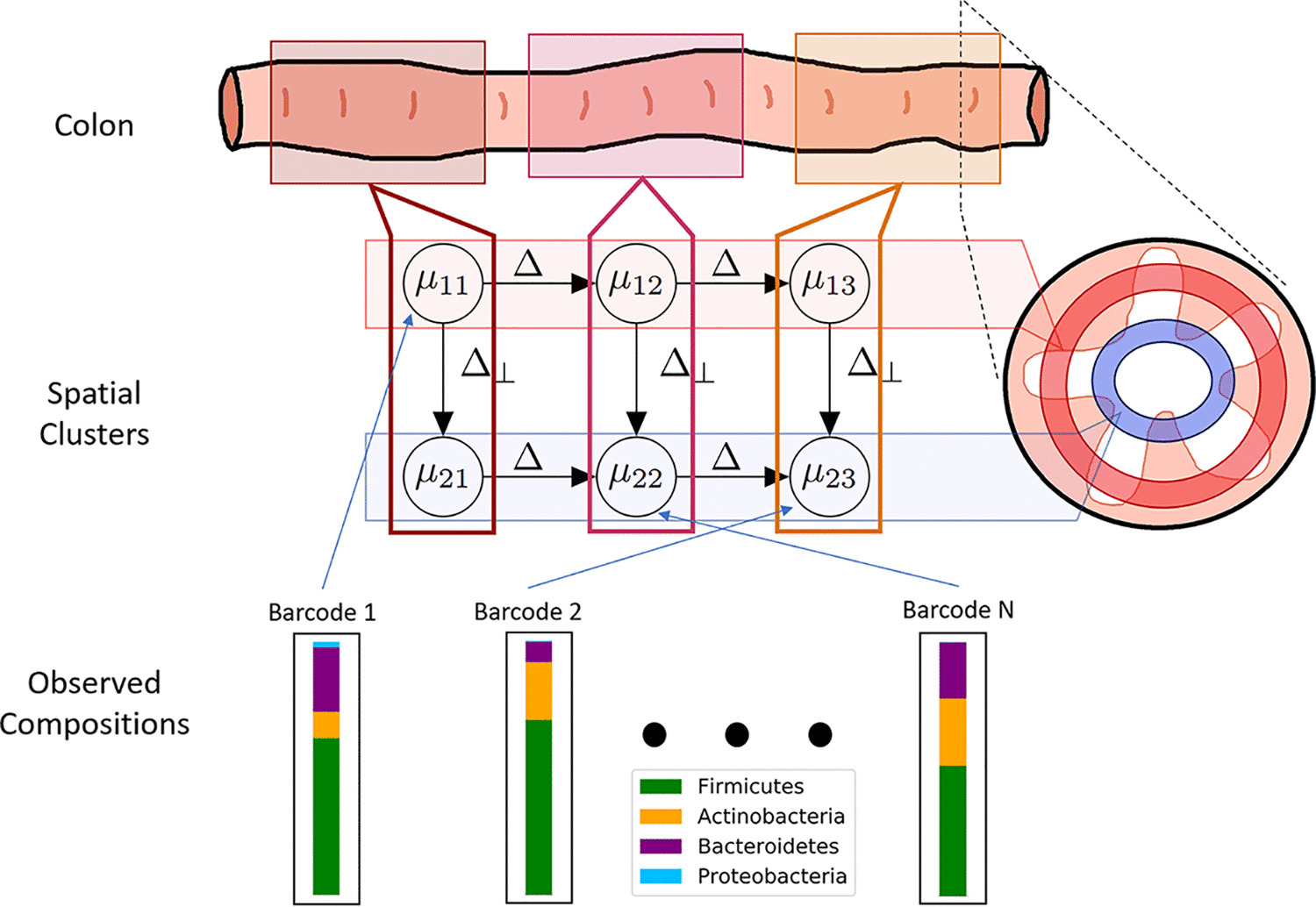
Directional Gaussian Mixture Models of the Gut Microbiome Elucidate Microbial Spatial Structure
Amey P. Pasarkar, Tyler A. Joseph, Itsik Pe'er mSystems, 2021 We develop a class of directional Gaussian Mixture models to elucidate the spatial arrangement of the gut microbiome. With no prior information, these models are able to recapitulate known spatial dynamics and propose new ones as well. Research was also presented at Probabalistic Modeling in Genomics, 2021. |
|
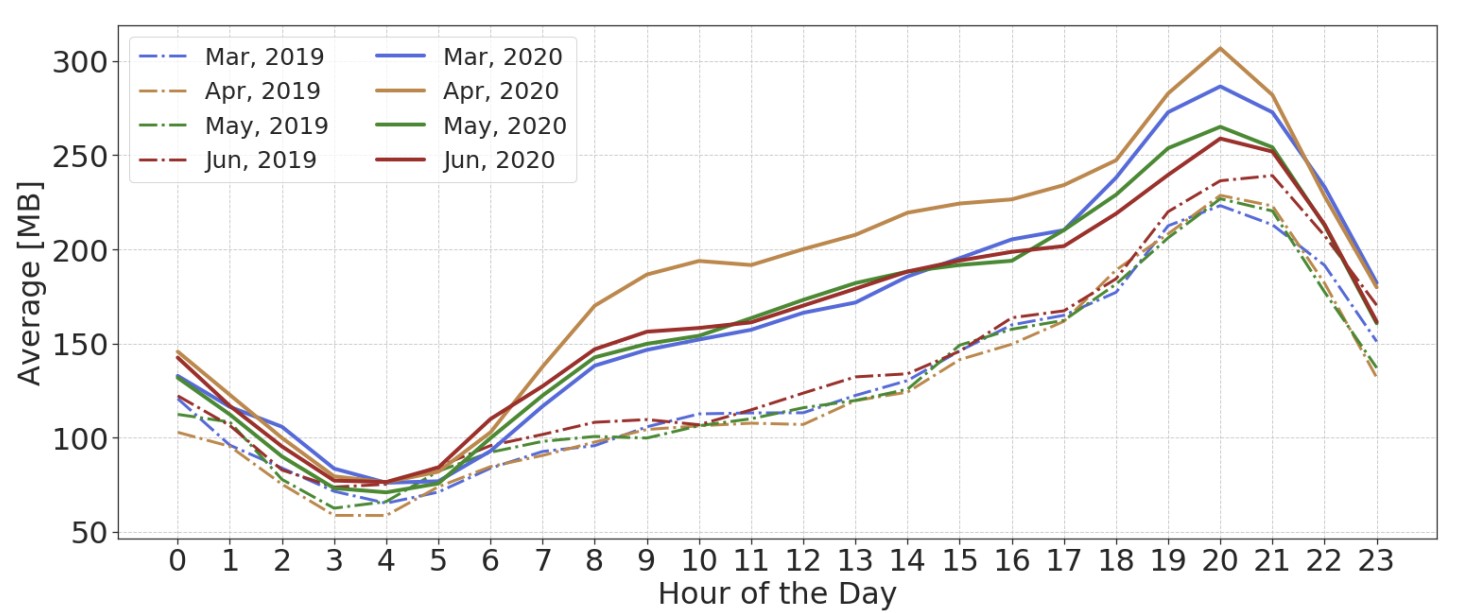
Social Distancing and the Internet: What Can Network Performance Measurements Tell Us?
Jessica De Oliveira Moreira, Amey P. Pasarkar, Wenjun Chen Kerry Hu, Jan Janak Henning Schulzrinne TPRC, 2021 We analyzed fixed broadband usage across the United States in the months following COVID-19 lockdowns to determine the effect of lockdowns on internet usage patterns and network performance. We founds signs of increased congestion following significant changes in user behavior. |
|
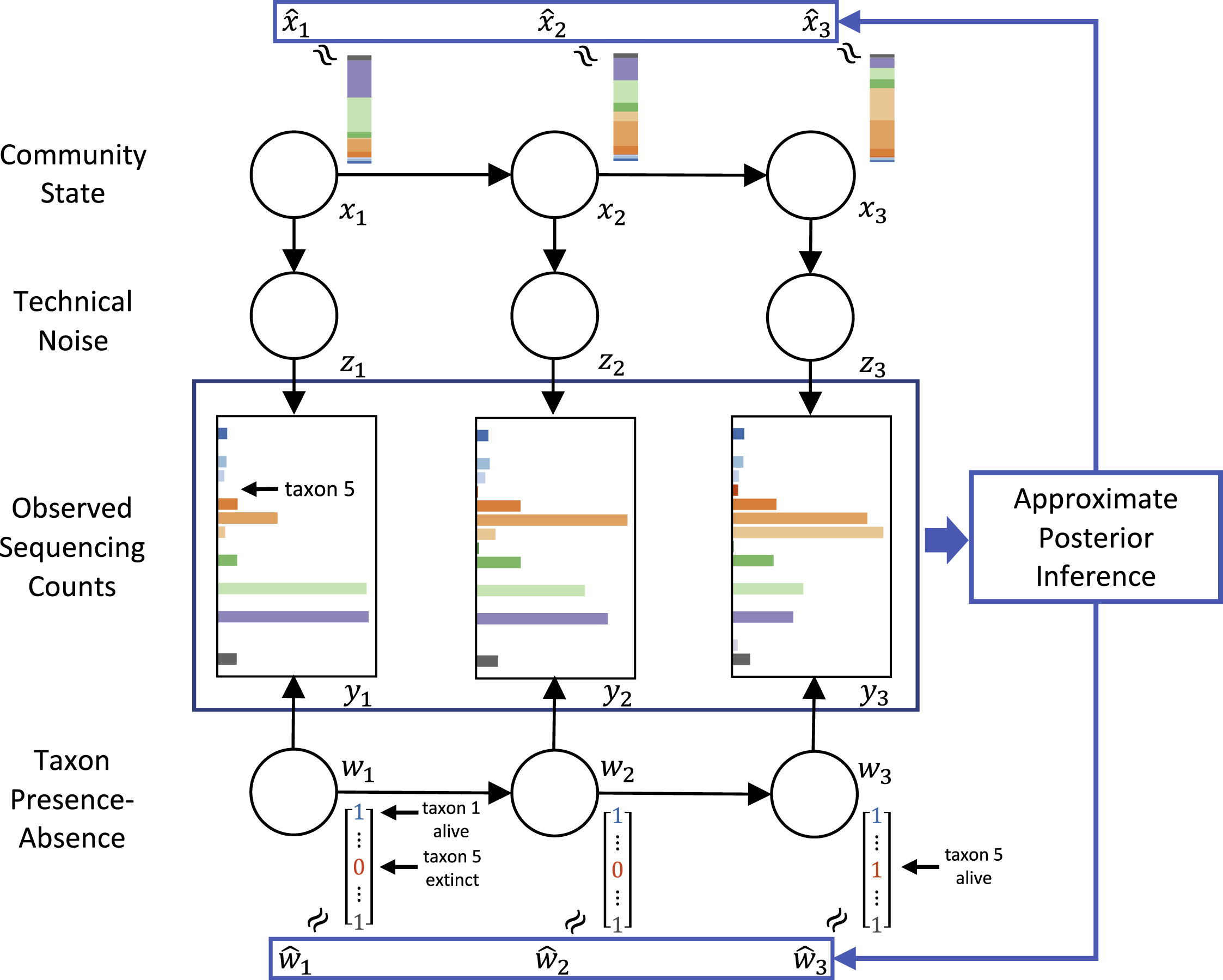
Efficient and Accurate Inference of Mixed Microbial Population Trajectories from Longitudinal Count Data
Tyler A. Joseph, Amey P. Pasarkar, Itsik Pe'er Cell Systems, 2020 We develop a multi-layer probabilistic graphical model that can deconvolve microbiome time-series data into its sources of noise. We show our model can be efficiently fit with variational inference while providing more accurate resuls than existing methods. Also presented at RECOMB 2020. |